By the Bioscience Core Lab team and Caitlin Clark
In the ongoing battle to learn more about SARS-CoV-2, the virus responsible for the COVID-19 pandemic, researchers from the KAUST Core Labs are contributing their expertise to help the world understand it.
The University’s Bioscience Core Lab (BCL) team recently sequenced the SARS-CoV-2 genome from 14 patient samples using Oxford Nanopore Technologies (ONT), a rapid, cost-effective and cutting-edge third generation sequencing (TGS) technology.
The samples came from a partnership with Arnab Pain, KAUST professor of bioscience, and his Pathogen Genomics lab group, who obtained them through his collaborations with local hospitals under the Saudi Ministry of Health.
To sequence the novel virus with ONT, the BCL team tailored the laboratory protocol through revisions of its chemistry and adapting the latest scientific advancements in the field.
BCL is the only ONT service provider in the Kingdom. The advanced technology’s use for SARS-CoV-2 represents important progress in the field of genomics in Saudi Arabia and is a key step in fighting the virus. It also holds promise for the University to collaborate with other research institutes and healthcare organizations in Saudi Arabia in the future.
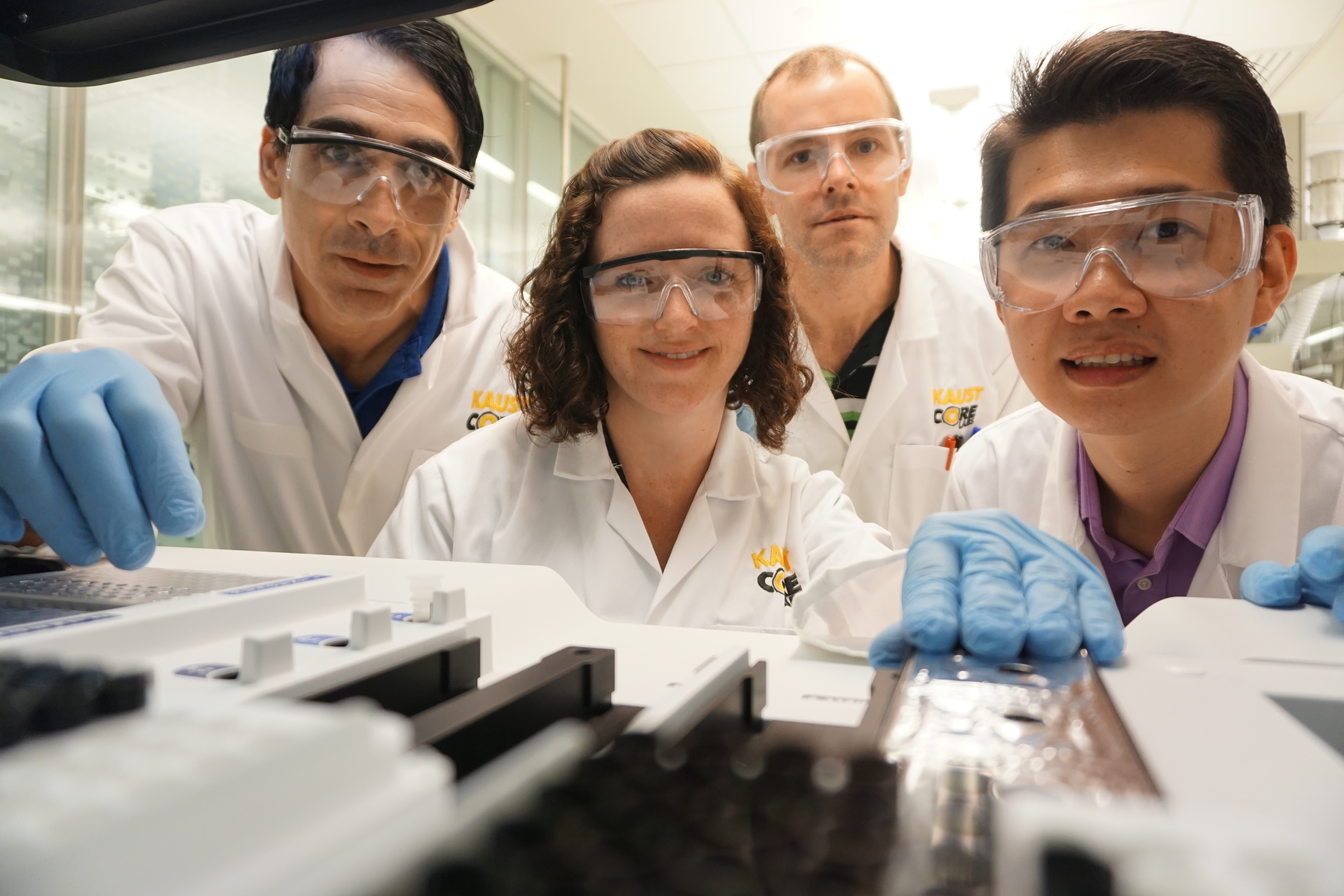

Bioscience Core Lab team members (from left to right) Luca Ermini, Karen Carty, Patrick Driguez and Alexander Putra make use of third generation sequencing technologies from Oxford Nanopore Technologies in the KAUST lab. Photo by Basil Chew.
Cutting-edge sequencing
ONT uses a nanopore—a protein with integrated sensors that is attached to a synthetic membrane—to sequence nucleic acids, which can be either DNA or RNA. The nanopore makes a nanometer-sized hole in the membrane, allowing a single strand of DNA or RNA to pass through.
A membrane full of nanopores is located on a flow cell, which is approximately the size of a glass microscope slide, and it is loaded with genomic libraries. The libraries hold genomic fragments with attached oligonucleotide adaptors and a motor protein that allows the genomic libraries to approach the nanopores. They are then pulled through for sequencing. The technology is able to operate hundreds to thousands of nanopores simultaneously.
“When the DNA or RNA passes through the nanopore, each nucleotide base making up the strand creates a different electrical signal that is then decoded and interpreted computationally to generate the sequence representation of DNA or RNA,” explained Luca Ermini, BCL team lead, Sanger and TGS.
“Compared to other older sequencing technologies, such as Sanger and next-generation sequencing, nanopore sequencing is a rapid, scalable and unique technology that enables direct real-time whole genome analysis,” Ermini continued. “The sequencing can be monitored live, which means that scientists know immediately if the process is going well, and it can be stopped at any time once there is enough data. For SARS-CoV-2 sequencing, we are able to generate enough depth coverage in approximately two hours.”

Luca Ermini (left), Bioscience Core Lab team lead, Sanger and third generation sequencing, uses the advanced GridION sequencer in the lab with a colleague. Photo by Andrea Bachofen-Echt.
Outstanding technology on campus
BCL uses several ONT platforms—including MinION, GridION and PromethION—for genome analysis, with all three allowing real-time data access.
“In early 2017, BCL began offering the MinION, a portable, handheld device that sequences one flow cell at a time,” noted Karen Carty, BCL senior technical specialist, TGS. “By late 2017, GridION, a benchtop sequencer that processes five flow cells in parallel, became available. In 2018, the PromethION, a second benchtop sequencer that is capable of high throughput sequencing, arrived on campus.”
“The GridION was used to sequence the first 14 samples of COVID-19 on two flow cells,” Ermini said. “However, as the SARS-CoV-2 genome is small (around 30 kilobases), 12 or 24—and potentially more—samples can be sequenced simultaneously per flow cell, allowing for further expanded use of the technology.”

KAUST Professor of Bioscience Arnab Pain (pictured) and his University research group are important collaborators for the Bioscience Core Lab team in sequencing SARS-CoV-2. Photo by Helmy Alsagaff.
Collaboration and analysis
Pain and his group carried out the extraction of viral RNA for library preparation, and they continue to analyze the sequenced samples to shed light on the biology of the virus.
“After extraction, the BCL team carried out a number of analytical steps, finally selecting and amplifying viral DNA with primers that span the entire SARS-CoV-2 genome,” explained Alexander Putra, BCL senior technical specialist, TGS. “After cleaning up the amplified DNA, barcoded adapters were added to distinguish between samples. Then, after loading to the flow cell, sequencing began immediately. Reads were then mapped—or compared—to the viral reference genome in real time to assess the quality of sequencing.”

Alexander Putra, Bioscience Core Lab (BCL) senior technical specialist, third generation sequencing, helped the BCL team modify the laboratory protocol to carry out sequencing of SARS-CoV-2. Photo by Andrea Bachofen-Echt.
“Following the completion of sequencing, individual base differences called SNPs and other changes in the genome, such as small deletions in the nucleotide sequences in the genome, were next determined and confirmed,” Pain said. “This is crucial for tracking changes in the virus populations as they pass through human populations and mutate—some of those mutations may be of important relevance to the disease progression and virulence.”
“The resulting viral genomes were then submitted by us to an open access online repository for a global comparison of virus mutations over time,” he continued. “Different mutations on different strains can be used to predict possible vaccine escape potential and also to identify regions in the genome of the virus that are under selection pressures. This information is important in designing potential vaccines.”
“Throughout, the sequencing process is monitored for quality in real time based on a bioinformatics protocol previously developed and validated by the scientific community,” noted Salim Bougouffa, BCL bioinformatics staff scientist. “In addition, the protocol and technology utilized have been used globally to study other outbreaks, such as those caused by Ebola, Zika and malaria.”
Scaling up for the future
“In the immediate future, we anticipate the possibility of sequencing more patient samples with the current protocol,” said Nicole Cheung, BCL director. “The data generated can be used to understand the genomic epidemiology and evolution of the virus and aid in vaccine development. This has a huge potential benefit for the Kingdom and the world.”
In addition, BCL team members note that the sequencing methods could be scaled up for use in any hospital lab in Saudi Arabia.
“The only things needed would be basic lab equipment, the necessary reagents, a portable MinION sequencer and laptop and the suitable protocol,” stated Patrick Driguez, BCL staff scientist, TGS. “Since it’s likely we’ll be combating the virus for an extended time period, on-site sequencing could help with tracking local outbreaks, determining their source and providing more data for the global scientific community.”

Bioscience Core Lab (BCL) scientist Luca Ermini noted that the BCL researchers will work on additional methods to study SARS-CoV-2 in the future. File photo.
“Further down the track, we will research additional methods useful for studying SARS-CoV-2,” continued Ermini. “Direct RNA sequencing is an obvious example—it can provide insights into the transcriptome and base modifications of the virus. Another approach could be metagenomics, which allows scientists to study other pathogens or microorganisms present in patient samples with the virus.”
“BCL sequencing the SARS-CoV-2 genomes with ONT is a milestone in Saudi sequencing history,” said Justin Mynar, KAUST associate vice president for research and executive director of the KAUST Core Labs and Research Infrastructure. “BCL’s expertise in cutting-edge sequencing technologies has the potential to make a real impact in the Kingdom and worldwide in the fight against COVID-19.”